Harder, Better, Faster:
Case Studies in Reproducible Workflows
Kathryn Huff
NYU Reproducibility Symposium
May 03, 2016
May 03, 2016









- Case Study Book Concept
- Case Study Contributions
- Lessons Learned
- Next Steps!


Reproducibility and Open Science Conference
May 21&22, 2015
- Three days
- Invitation Only
- Case Studies, Education, Self-assessment
- https://github.com/BIDS/repro-conf
Editors



Justin Kitzes, Fatma Imamoglu, Daniel Turek
Supplementary Chapter Authors

|

|

|
|
|
Fernando Chirigati |
Case Study Chapter Contributors!
|
|
|
|
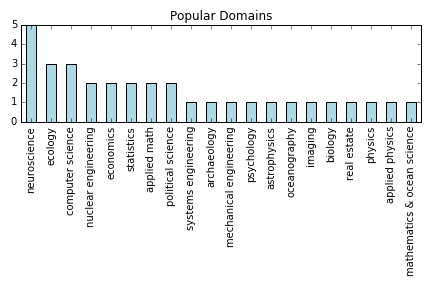
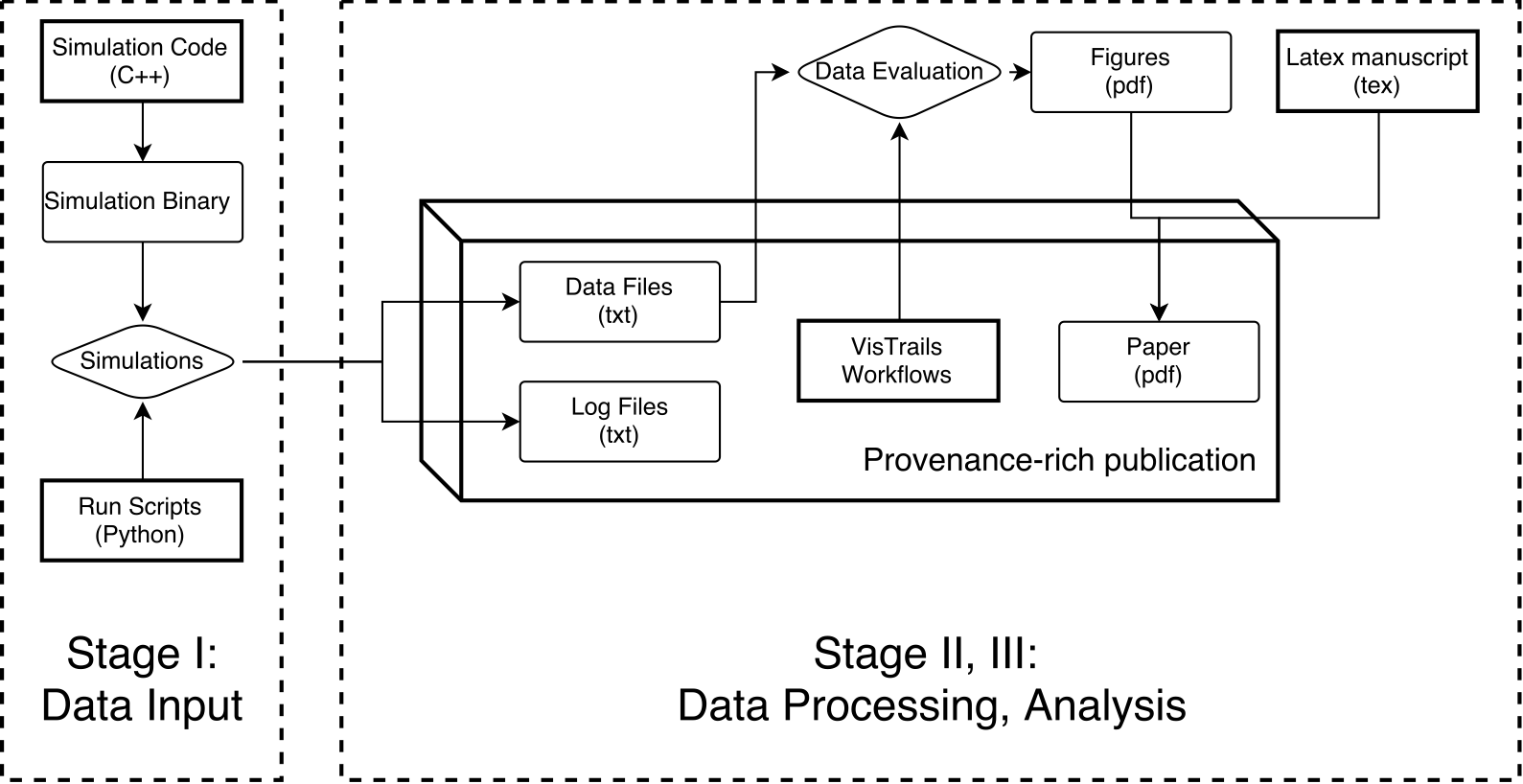
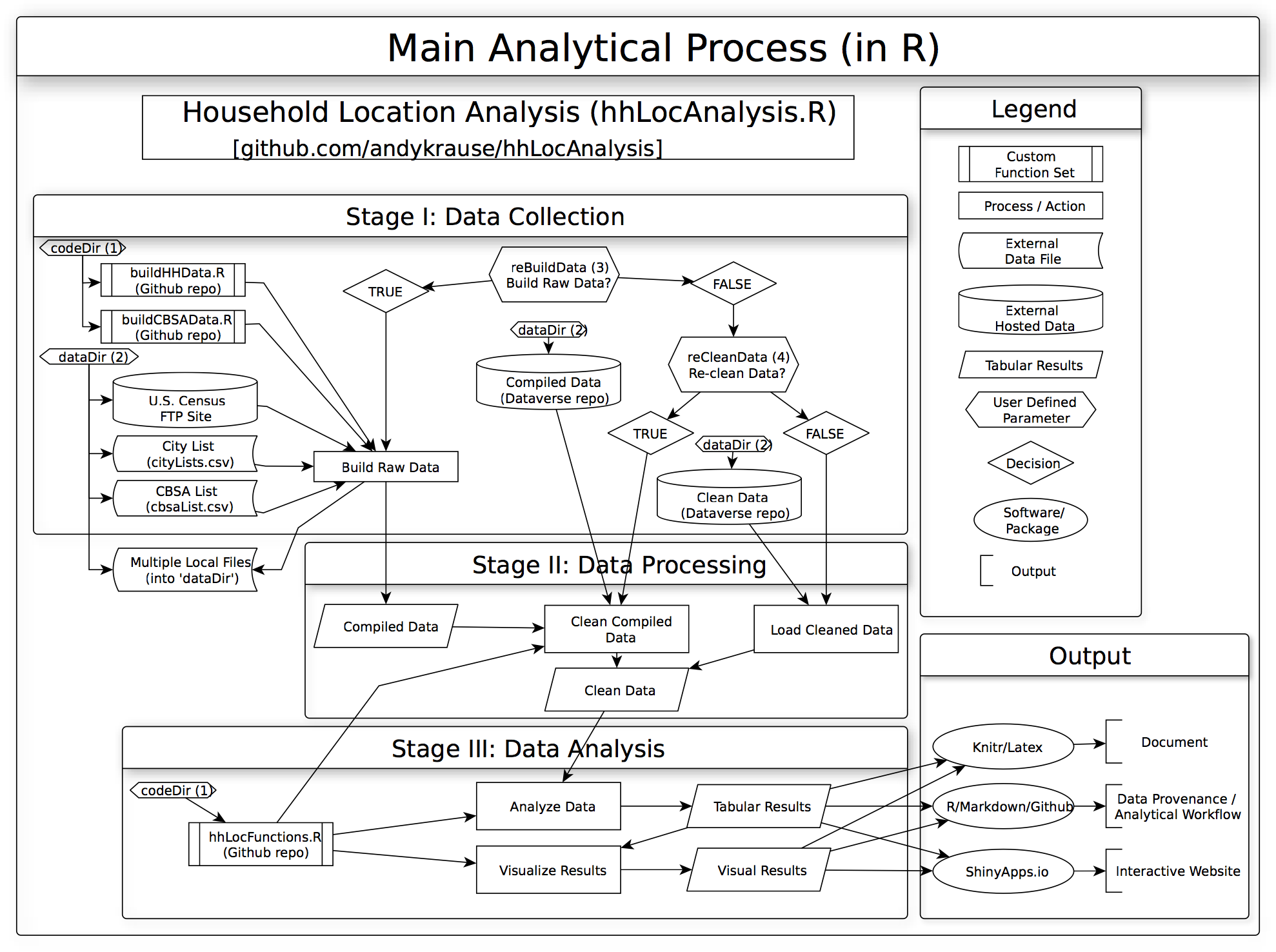
Pain Points
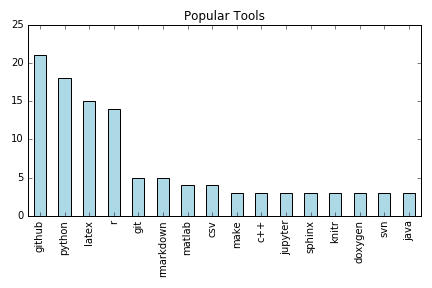
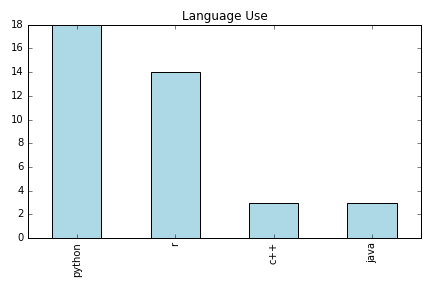
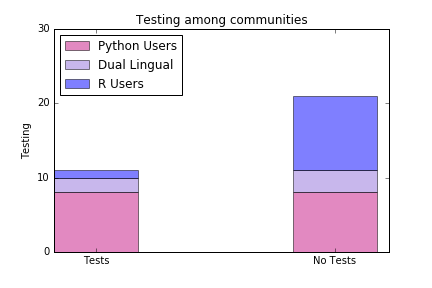
Emergent Needs
- Better education of scientists in more reproducibility-robust tools.
- Widely used tools should be more reproducible so that the common denominator tool does not undermine reproducibility.
- Improved configuration and build systems for portably packaging software, data, and analysis workflows.
- Reproducibility at scale for high performance computing.
- Standardized hardware configurations and experimental procedures for limited-availability experimental apparatuses.
- Better understanding of why researchers don't respond to the delayed incentives of unit testing as a practice.
- Greater adoption of unit testing irrespective of programming language.
- Broader community adoption around publication formats that allow parallel editing (i.e. any plain text markup language that can be version
- Greater scientific adoption of new industry-led tools and platforms for data storage, versioning, and management.
- Increased community recognition of the benefits of reproducibility.
- Incentive systems where reproducibility is not self-incentivizing.
- Standards around scrubbed and representational data so that analysis can be investigated separate from restricted data sets.
- Community adoption for file format standards within some domains.
- Domain standards which translate well outside of their own scientific communities.
Social Science Volume
Collecting Case Studies Spring/Summer 2016
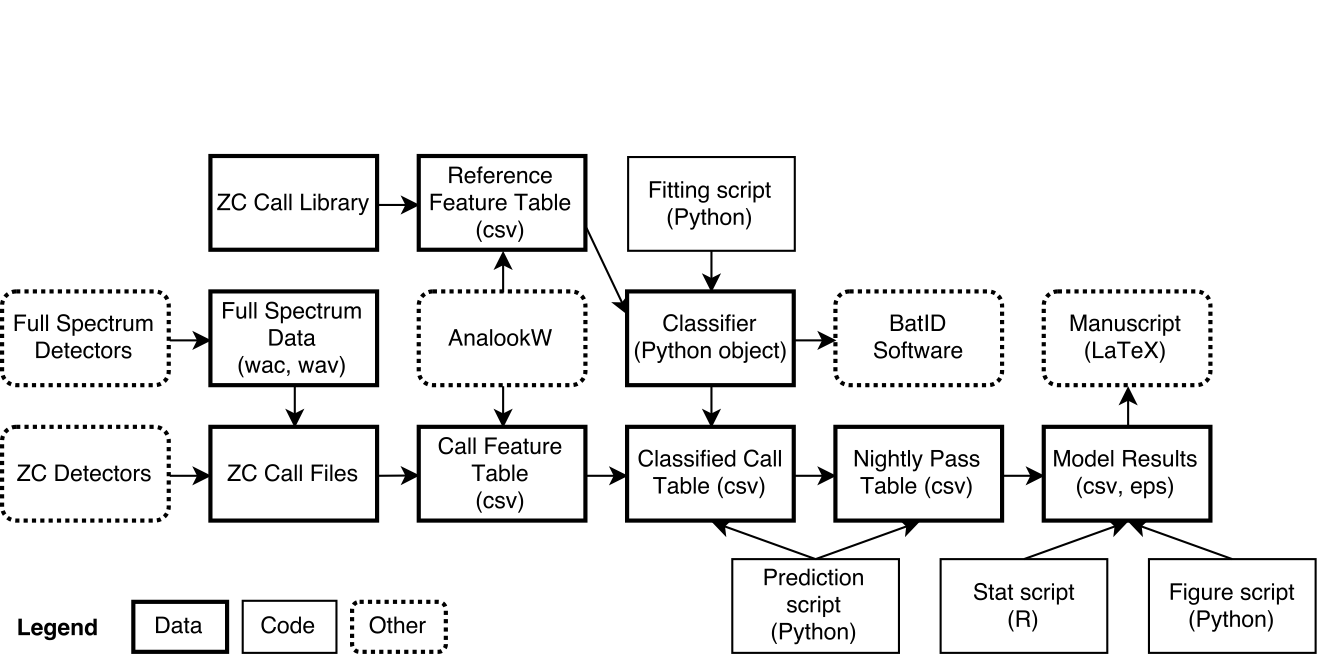
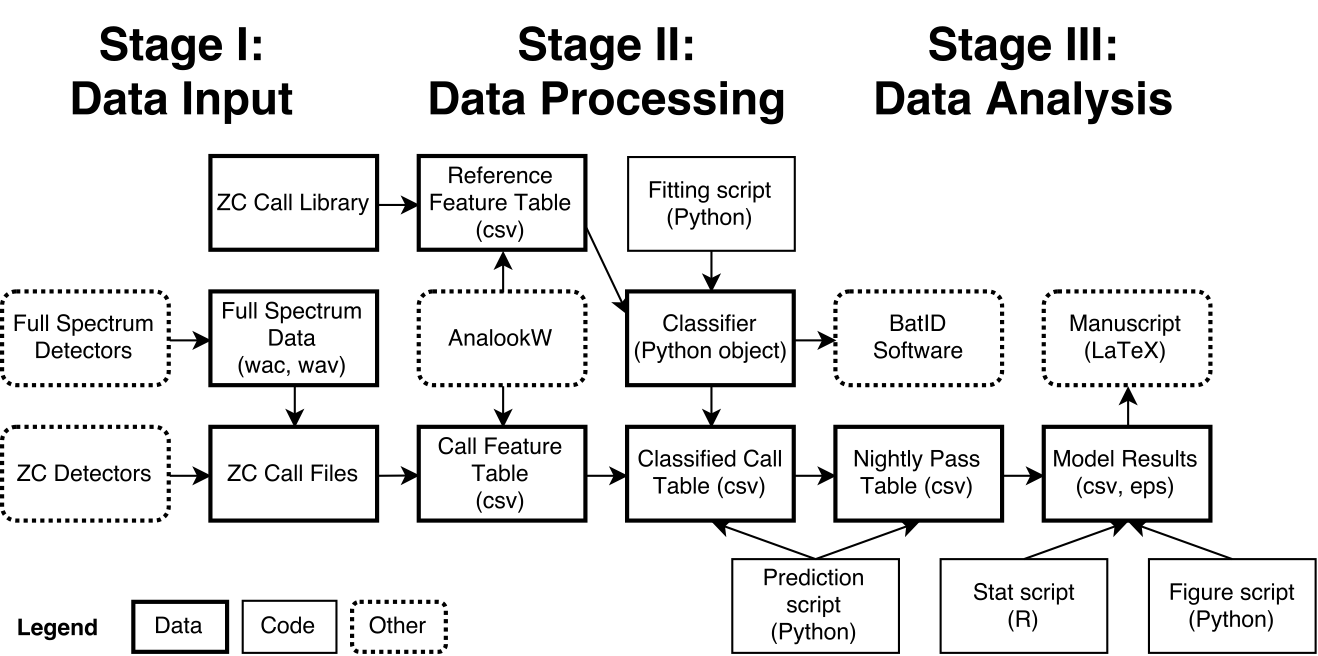
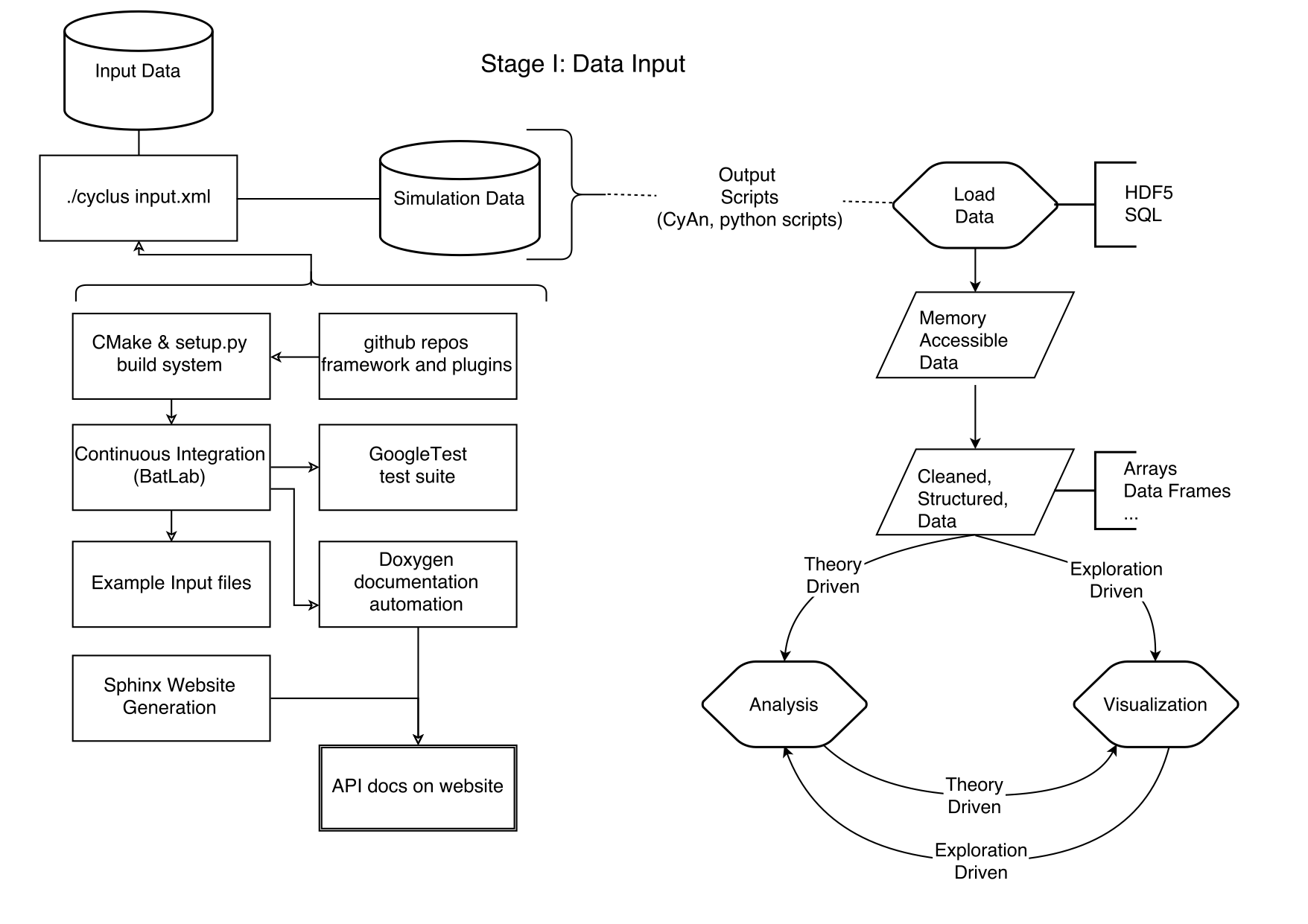
- Same format: 1,500-2,000 words plus one diagram
- Bad Hessian blog : http://www.badhessian.org
- GitHub Repo : http://github.com/BIDS/ss-repro-case-public
- Email Garret Christensen (garret@berkeley.edu) or Cyrus Dioun (dioun@berkeley.edu)
Acknowledgements
- Justin Kitzes
- Fatma Imamoglu
- Daniel Turek
- Chapter Authors
- Case Study Authors
- Reproducibility Working Group

THE END
Katy Huff
katyhuff.github.io/2016-05-03-nyu
Harder, Better, Faster: Case Studies in Reproducible Workflows by Kathryn Huff is licensed under a Creative Commons Attribution 4.0 International License.
Based on a work at http://katyhuff.github.io/2016-05-03-nyu.